How to Prepare Data for BESA Connectivity
| Module information | |
| Modules | BESA Research Basic or higher,
BESA Connectivity |
| Version | 7.0 or higher (BR), 1.0 or higher (BC) |
Contents
Note and general remarks
This article applies to data that are prepared using BESA Research. It is also possible to read data prepared by other programs. For the requirements, please refer to the BESA Connectivity manual, the chapter on "Loading data that were created in other software" and the chapter "File formats".
BESA Connectivity needs the following data:
- A file containing the binary data matrix. All epochs in the data file need to have the same length, the same stimulus position, the same baseline interval, and the same channel configuration.
- A header file in ASCII format.
- A channel description file in ASCII format. Two types of files are supported: elp files and bsa files.
File format descriptions
Header file format
The header file is a standard ASCII file. It contains the following lines:
| Line | Entry | Possible values | Description |
|---|---|---|---|
| 1 | BESA Generic Data v1.1 | File format version | |
| 2 | nChannels= | [1 ... 1024] | Number of channels. |
| 3 | sRate= | [0.0001 ... ] | Sampling rate (samples/second) in Hz. |
| 4 | nSamples= | [1 ...] | Total number of samples in file. |
| 5 | format= | float | Format of the data in the binary file. |
| 6 | file= | Name of binary file. | Name of binary file without path information. |
| 7 | prestimulus= | 500.000 | Pre-stimulus interval in milliseconds. If negative, then the epoch starts after the stimulus. This effectively means that the value of prestimulus denotes the position of the stimulus in the epoch (milliseconds after epoch start). |
| 8 | epochs= | Number of epochs in the data file. | |
| 9 | baselineStart= | Baseline start relative to the stimulus position. The value should be given in milliseconds. | |
| 10 | baselineEnd= | Baseline end relative to the stimulus position. Must be >= baselineStart. The value should be given in milliseconds. | |
| 11 | epochLength= | Epoch length in milliseconds. This parameter is optional and can be added for convenience. If not given, the epoch length is calculated from the overall number of samples, the sampling rate, and the number of epochs. | |
| 11 | Padding= | 2000.000 | Information concerning extra data values that surround the epoch to provide padding for wavelets, expressed in milliseconds. If exported from BESA Research the default value is 2000ms. |
| 12 | conditionName= | Name of the condition that identifies it. | |
| 13 | channelUnits= | Unit that the data are stored in the binary file. For EEG data, it should be one of µV, mV, V, µV/cm². For source data, it should be one of nAm, nAm/cm², nAm/cm³. For MEG data, it should be one of fT, pT, fT/cm, pT/cm, µV/cm².
This entry is repeated <nChannels> times. |
Channel configuration file format
The channel description file is an ASCII file. Two types of files are supported: .elp files and .bsa files.
ELP file format
Elp files describe surface channel coordinates using spherical coordinates (theta, phi). It contains one line for each channel in the data file. Each line contains the following entries:
| Column | Entry | Possible values | Description |
|---|---|---|---|
| 1 | Channel type | EEG (scalp electrode) SCP (scalp electrode) |
This column denotes the type of the channel. |
| 2 | Channel label | Fpz Fz |
Channel label. Is not allowed to contain blanks. |
| 3 | Azimuth (theta) | [-180.00 ... 180.00] | Spherical azimuth angle in degrees (optional entry). |
| 4 | Latitude (phi) | [-90.00 ... 90.00] | Spherical latitude angle in degrees (optional entry). |
BSA file format
Bsa files describe source channel data. They have a header section for format version and readability, followed by one line for each source channel.
| Column | Entry | Possible values | Description |
|---|---|---|---|
| 1 | Channel type | RegSrc (regional source)
DipSrc (dipole) |
This column denotes the type of the channel. |
| 2 | x location | x coordinate of the channel. | |
| 3 | y location | y coordinate of the channel. | |
| 4 | z location | z coordinate of the channel. | |
| 5 | x orientation | x orientation of the channel. | |
| 6 | y orientation | y orientation of the channel. | |
| 7 | z orientation | z orientation of the channel. | |
| 8 | Color | [0 … 255*255*255] | RGB color value |
The coordinate system is defined by a sphere with unit dimensions, which is fitted to the coordinates of sensors on the head. In the absence of co-registration information with individual MRI data, the axes are defined by reference points on the head known as fiducials. The reference points are normally the nasion (NAS), the left preauricular point (LPA), and the right preauricular point (RPA). The direction of the x axis is defined by the line joining LPA and RPA, positive towards RPA. The direction of the y axis is defined by the line through NAS that is perpendicular to the x axis (positive towards NAS). The z axis is perpendicular to the x and y axes, and goes up out of the upper part of the head (e.g. Cz). On average, the center of the unit sphere is about 4 cm above the origin of the head coordinate system.
Binary data matrix file format
The binary data matrix is saved in float format and contains <Number of epochs> * <Number of samples per epoch> * <Number of channels> values. These are stored in the following order:
- Slowest index: epochs
- Next index: samples
- Fastest index: channels
This is the so-called multiplexed file format. The default extension for this header file is: *.dat
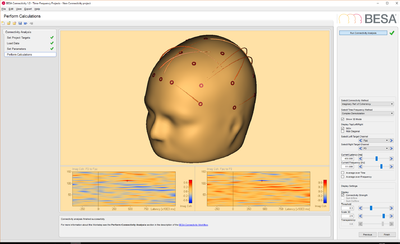
Export data using BESA Research
Export data using Matlab
Examples
Minimal examples for data files that can be imported in BESA Connectivity can be downloaded here.
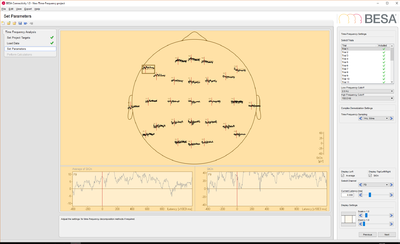
- EEG data: this data set contains 25 trials (trial length: -400ms to 1200ms) of scalp data with 27 electrodes at a sampling rate of 320 samples/second.
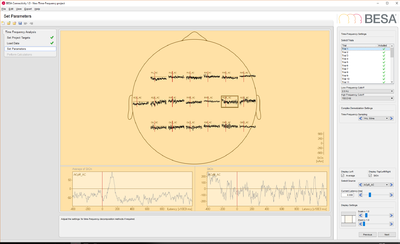
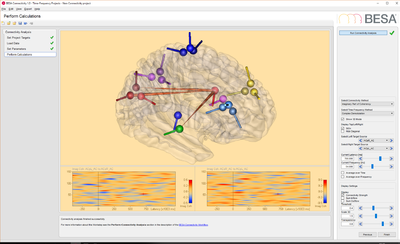
- Source data: this data set contains 33 trials (trial length: -400ms to 1200ms) of source data with 19 dipoles at a sampling rate of 320 samples/second.